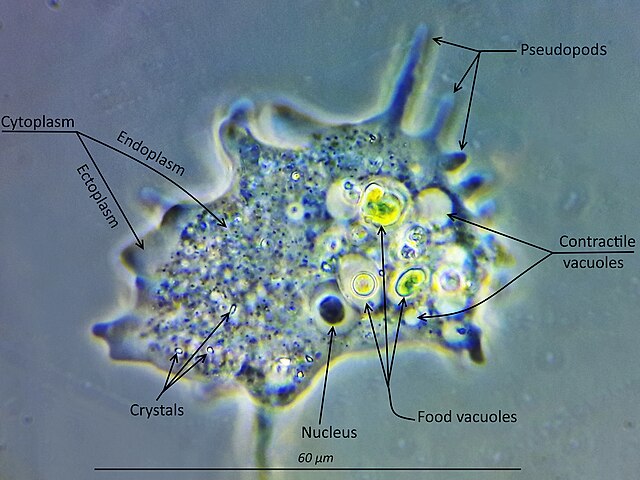
Entamoeba is a genus of Amoebozoa found as internal parasites or commensals of animals. In 1875, Fedor Lösch described the first proven case of amoebic dysentery in St. Petersburg, Russia. He referred to the amoeba he observed microscopically as Amoeba coli; however, it is not clear whether he was using this as a descriptive term or intended it as a formal taxonomic name. The genus Entamoeba was defined by Casagrandi and Barbagallo for the species Entamoeba coli, which is known to be a commensal organism. Lösch's organism was renamed Entamoeba histolytica by Fritz Schaudinn in 1903; he later died, in 1906, from a self-inflicted infection when studying this amoeba. For a time during the first half of the 20th century the entire genus Entamoeba was transferred to Endamoeba, a genus of amoebas infecting invertebrates about which little is known. This move was reversed by the International Commission on Zoological Nomenclature in the late 1950s, and Entamoeba has stayed 'stable' ever since.

Entamoeba
Entamoeba gingivalis
Amoebozoa is a major taxonomic group containing about 2,400 described species of amoeboid protists, often possessing blunt, fingerlike, lobose pseudopods and tubular mitochondrial cristae. In traditional classification schemes, Amoebozoa is usually ranked as a phylum within either the kingdom Protista or the kingdom Protozoa. In the classification favored by the International Society of Protistologists, it is retained as an unranked "supergroup" within Eukaryota. Molecular genetic analysis supports Amoebozoa as a monophyletic clade. Modern studies of eukaryotic phylogenetic trees identify it as the sister group to Opisthokonta, another major clade which contains both fungi and animals as well as several other clades comprising some 300 species of unicellular eukaryotes. Amoebozoa and Opisthokonta are sometimes grouped together in a high-level taxon, variously named Unikonta, Amorphea or Opimoda.
Amoebozoa
An amoeba of the genus Mayorella (Amoebozoa, Discosea)
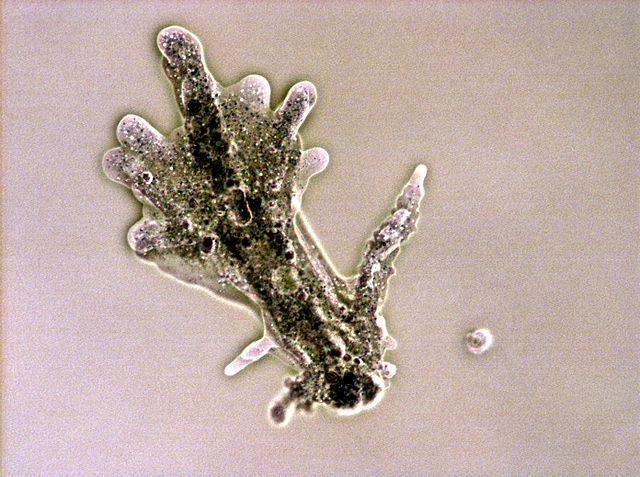
Amoeba proteus (Lobosa: Tubulinea)
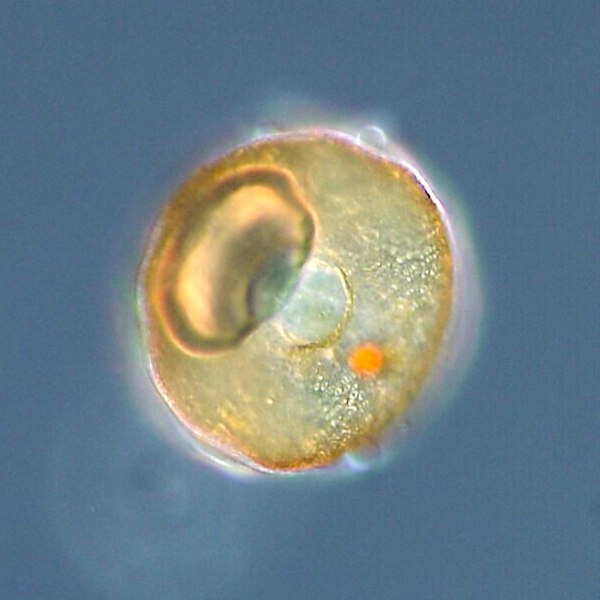
Arcella sp. test (Lobosa: Tubulinea)