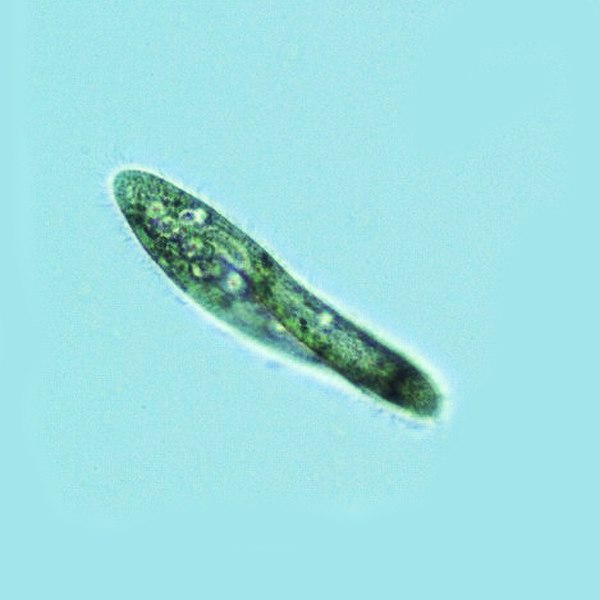
Tetrahymena, a unicellular eukaryote, is a genus of free-living ciliates. The genus Tetrahymena is the most widely studied member of its phylum. It can produce, store and react with different types of hormones. Tetrahymena cells can recognize both related and hostile cells.
Tetrahymena conjugation. When nutrients are scarce, two individuals (A) pair with each other and begin sexual reproduction (conjugation). (B) The diploid micronucleus in each individual undergoes meiosis to form four haploid nuclei, and three of these are degraded. (C) The remaining haploid nucleus divides mitotically to form two pronuclei in each cell. (D) One of the two pronuclei in each cell is exchanged with the mating partner, and fusion leads to the formation of the diploid
Two Tetrahymena cells of complementary mating types pair to exchange nuclei during sexual conjugation.
A unicellular organism, also known as a single-celled organism, is an organism that consists of a single cell, unlike a multicellular organism that consists of multiple cells. Organisms fall into two general categories: prokaryotic organisms and eukaryotic organisms. Most prokaryotes are unicellular and are classified into bacteria and archaea. Many eukaryotes are multicellular, but some are unicellular such as protozoa, unicellular algae, and unicellular fungi. Unicellular organisms are thought to be the oldest form of life, with early protocells possibly emerging 3.8–4.8 billion years ago.
Valonia ventricosa, a species of alga with a diameter that ranges typically from 1 to 4 centimetres (0.4 to 1.6 in) is among the largest unicellular species
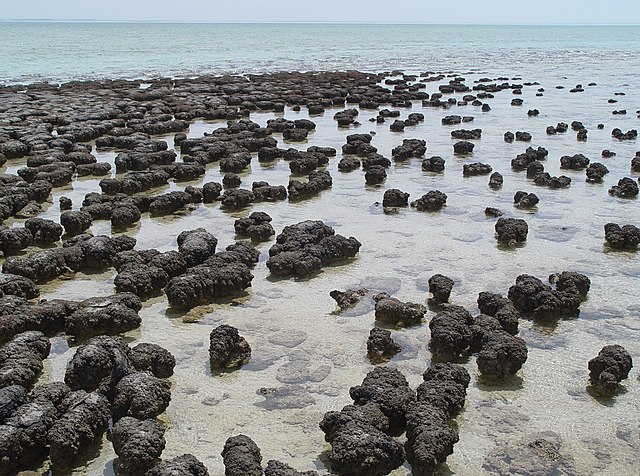
Modern stromatolites in Shark Bay, Western Australia. It can take a century for a stromatolite to grow 5 cm.
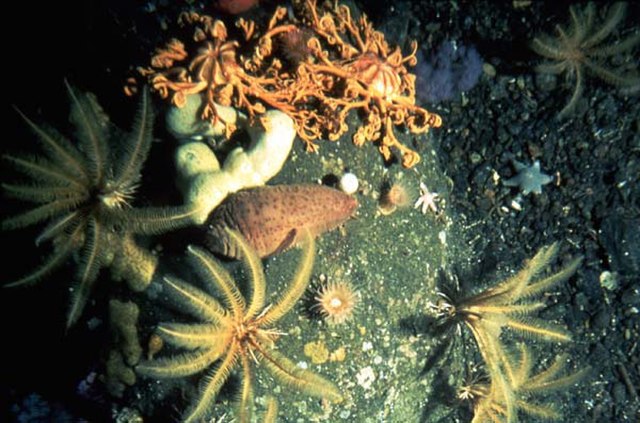
A bottom-dwelling community found deep in the European Arctic.
Paramecium tetraurelia, a ciliate, with oral groove visible