An Alu element is a short stretch of DNA originally characterized by the action of the Arthrobacter luteus (Alu) restriction endonuclease. Alu elements are the most abundant transposable elements in the human genome, present in excess of one million copies. Alu elements were thought to be selfish or parasitic DNA, because their sole known function is self reproduction. However, they are likely to play a role in evolution and have been used as genetic markers. They are derived from the small cytoplasmic 7SL RNA, a component of the signal recognition particle. Alu elements are highly conserved within primate genomes and originated in the genome of an ancestor of Supraprimates.
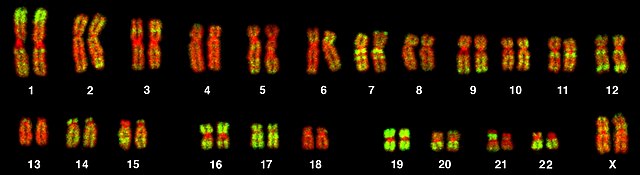
Karyotype from a female human lymphocyte (46, XX). Chromosomes were hybridized with a probe for Alu elements (green) and counterstained with TOPRO-3 (red). Alu elements were used as a marker for chromosomes and chromosome bands rich in genes.
The human genome is a complete set of nucleic acid sequences for humans, encoded as DNA within the 23 chromosome pairs in cell nuclei and in a small DNA molecule found within individual mitochondria. These are usually treated separately as the nuclear genome and the mitochondrial genome. Human genomes include both protein-coding DNA sequences and various types of DNA that does not encode proteins. The latter is a diverse category that includes DNA coding for non-translated RNA, such as that for ribosomal RNA, transfer RNA, ribozymes, small nuclear RNAs, and several types of regulatory RNAs. It also includes promoters and their associated gene-regulatory elements, DNA playing structural and replicatory roles, such as scaffolding regions, telomeres, centromeres, and origins of replication, plus large numbers of transposable elements, inserted viral DNA, non-functional pseudogenes and simple, highly repetitive sequences. Introns make up a large percentage of non-coding DNA. Some of this non-coding DNA is non-functional junk DNA, such as pseudogenes, but there is no firm consensus on the total amount of junk DNA.

TSC SNP distribution along the long arm of chromosome 22 (from https://web.archive.org/web/20130903043223/http://snp.cshl.org/ ). Each column represents a 1 Mb interval; the approximate cytogenetic position is given on the x-axis. Clear peaks and troughs of SNP density can be seen, possibly reflecting different rates of mutation, recombination and selection.
Populations with a high level of parental-relatedness result in a larger number of homozygous gene knockouts as compared to outbred populations.
A pedigree displaying a first-cousin mating (carriers both carrying heterozygous knockouts mating as marked by double line) leading to offspring possessing a homozygous gene knockout